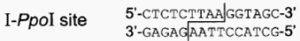Beware the new 'Breakthrough' Transgenic Mosquitoesby Dr Mae Wan Ho
6/24/14
NOTICE: THIS WORK MAY BE PROTECTED BY COPYRIGHTYOU ARE REQUIRED TO READ
THE COPYRIGHT NOTICE AT THIS LINK BEFORE YOU READ THE FOLLOWING WORK, THAT IS AVAILABLE SOLELY FOR PRIVATE STUDY, SCHOLARSHIP OR RESEARCH PURSUANT TO 17 U.S.C. SECTION 107 AND 108. IN THE EVENT THAT THE LIBRARY DETERMINES THAT UNLAWFUL COPYING OF THIS WORK HAS OCCURRED, THE LIBRARY HAS THE RIGHT TO BLOCK THE I.P. ADDRESS AT WHICH THE UNLAWFUL COPYING APPEARED TO HAVE OCCURRED. THANK YOU FOR RESPECTING THE RIGHTS OF COPYRIGHT OWNERS.
Mosquitoes engineered with a jumping gene vector to express a DNA-cutting enzyme produce >95 % male offspring; unfortunately both enzyme and vector target genomes of diverse species from slime moulds to humans. Dr Mae Wan Ho
This commentary has been sent to the editors of the journal Nature Communications, inviting them and the researchers who have reported the creation of the new transgenic mosquitoes in the journal to reply.
Please circulate widely and repost, but you must give the URL of the original and preserve all the links back to articles on our website. If you find this report useful, please support ISIS by subscribing to our magazine Science in Society, and encourage your friends to do so. Or have a look at the ISIS bookstore for other publicationsA good trick but no consideration of risksA team led by Andrea Crisanti at Imperial College London in the UK was widely reported to have made a breakthrough or even a ‘quantum leap’ in creating transgenic mosquitoes that could eradicate malaria [1]. Unfortunately, it is potentially the most hazardous genetically modified organism (GMO) to have been created, and should go no further from the laboratory. The researchers have not considered the risks involved, which would have been obvious from a casual review of existing literature.Their fast-tracked online report in Nature Communications stated [2]: “Here we generate a synthetic sex distortion system by exploiting the specificity of the homing endonuclease I-PpoI, which is able to selectively cleave ribosomal gene sequences of the malaria vector Anopheles gambiae that are located exclusively on the mosquito’s X-chromosome. We combine structure-based protein engineering and molecular genetics to restrict the activity of the potentially toxic endonuclease to spermatogenesis. Shredding of the paternal X-chromosome prevents it from being transmitted to the next generation resulting in fully fertile mosquito strains that produce >95 % male offspring.”
Simply considered as a genetic trick, it is ingenious. Shredding the X chromosome of the male will make all of its offspring males. That is because female mosquitoes (like female humans) have two X chromosomes, one from the male parent and the other from the female parent, so without the contribution of the X chromosome from the male parent, only male offspring will result. A completely sterile male mosquito is useless, as it just dies out without affecting the population. But a fully fertile one that breeds exclusively males and pass on the sex-distorter trait would be ideal, as it would indeed wipe out the natural population, provided the trait is stably inherited. It would have been the perfect solution to destroying the natural populations of mosquitoes that transmit malaria; except that the DNA-cutting enzyme is by no means “specific” to “ribosomal gene sequences located exclusively on the mosquito’s X-chromosome” as stated. On the contrary, it cuts at a target sequence in ribosomal RNA (rRNA) genes -- numerous copies of which are present in all eukaryote genomes -– plus other sites as well, and the transgenic mosquitoes have been created using a jumping gene (transposon) vector that promiscuously invades all genomes. It is the female mosquitoes that bite people and transmit disease; so any transgenic female mosquitoes among the offspring would inject GM DNA containing the vector and I-PpoI transgene for horizontal transfer into people’s cells to shred their genomes.The creation of transgene mosquitoes with heritable sex distortionThe wild-type I-PpoI has been engineered into mosquitoes. The expression of the wild-type enzyme during spermatogenesis in transgenic mosquitoes causes cleavage of the paternal X chromosome, but also results in complete male sterility because the protein is stable and persists in mature sperm cells, leading to subsequent cleavage of the maternal X chromosome in the zygote after fertilization, thereby killing the zygote. Thus, the sterile males would just die out without leaving any offspring, and not make any difference to the natural population, unless they are continually released into the wild.
To overcome that, the team mutated amino acid residues in the hydrophobic core of the endonuclease to obtain recombinant proteins with reduced stability. The melting temperature of the protein was decreased from 54.4 °C in the wild type to 49.4 °C in mutant L111A and 35.1 °c in a double mutant L111A/W124L. The thermal half-life at 37 °C ranged from 73.5h for the wild type enzyme to 2 h for H106A. The L111A/W 124L double mutant had a half-life of about 4 min. The specific activity of the wild type was ~7 pmol/min/mg compared to about a third of that in mutant H106A.
Germ line transformation constructs were created to express the I-PpoI variants placed under the control of the male spermatogenesis-specific b2-tubulin promoter. The constructs were designed to express both enhanced green fluorescent protein (EGFP) as in frame fusion protein or, using a 2A ribosomal stuttering signal, as distinct protein chains. The constructs were put into the promiscuous piggyBac vectors for transgenesis. Constructs that had integrated on the X chromosome failed to show significant levels of I-PpoI expression in the testes, probably because the X chromosome is transcriptionally silent during male meiosis.
Reduced in vitro thermal stability translated into significantly lower protein levels of EGFPI-PpoI protein in vivo. The destabilized I-PpoI distorts the sex ratio towards males as measured in emerging adults, fertility was measured by larval hatching rate and number of eggs in crosses of transgenic male mosquitoes to wild type females. As a control transgenic female mosquitoes of each strain were crossed to wild type males. Male mosquitoes expressing the wild-type enzyme were sterile as previously found. No effect on fertility or sex ratio was observed in all strains expressing the double mutant L111A/W124L, which had the lowest in vitro stability; or in strains carrying X-linked transgenes. Significantly, male biased sex ratios ranging from 70.2 to 97.4 % were found in the progeny of males carrying the remaining I-PpoI variants; of those, only three lines, 111A-2 and 124L-2, 124L-3 had comparable hatching rates to the wild type. These strains also showed the lowest mRNA levels.
The sex distortion phenotype was stably inherited from male mosquitoes to their transgenic sons. For four subsequent generations, 111A-2 fathers showed comparable levels of fertility and male-biased sex ratios in their offspring. Homozygous males in strains 111A-2 and 124L-3 caused a reduction in the hatching rate, but had no effect on fertility in 124L-2.
It is important to note that the sex distortion is not complete, which means that a variable number of transgenic females will be left to bite people and transmit the potentially lethal transgenes into people’s cells. Such transgenic females were found to have more female offspring when mated to wild type males (see below). In the wild, the proportion of surviving transgenic females, and indeed, I-PpoI-resistant transgenic females could also arise (see below). All that would make it more likely to spread the transgenes into human and other mammals.Female offspring of transgenic male mosquitoes showed evidence of misrepair and copy number variation in the ribosomal gene cluster in their genome, suggesting chromosomal damage. There was also a significant female-biased sex ratio in the progeny of female survivors crossed to wild type males suggesting that loss of vitality occurred in individuals that had inherited only a damaged X chromosome from the transgenic female.
But have the transgenic female offspring inherited integrated I-PpoI endonuclease genes in the X-chromosome, which would make the chromosome resistant to further endonuclease damage (as integration destroys the target site, see below)? To test for this possibility, the transgenic females were crossed to wild type males, and the transgenic male progeny, which have a 50 % chance of carrying the potentially resistant X-chromosome was then crossed to wild type females, and the sex ratio of individual crosses was recorded.
The analysis showed a male bias in the progeny of all males that did not differ significantly from the 111A-2 stock, suggesting that the X chromosomes of female survivors are damaged but still susceptible to further cleavage.In five independent cage experiments, the release of hemizygous 111A-2 males at a ratio of 3X controls was effective in suppressing caged wild type populations (achieving elimination within 6 generations in four out of five cages. As expected the release of sterile males expressing the wild-type enzyme had no measurable effect in three populations. Sex distorter is much more effective than sterile males by 2 orders of magnitude, and 2-70 times more effective than equivalent releases of female-killing alleles.
This sex-distorter system is clearly an improvement on the Oxytec transgenic mosquitoes engineered with an ill-characterized, ineffective, as well as hazardous system [3, 4] (Regulation of Transgenic Insects Highly Inadequate and Unsafe, Transgenic Mosquitoes Not a Solution, SiS 54).
There is an alternative non-transgenic system based on a common symbiotic bacterium that can stop the dengue virus multiplying in the mosquito host, which effectively makes those transgenic mosquitoes obsolete [5] (Non-transgenic Mosquitoes to Combat Dengue, SiS 54).
But the ill-advised Brazilian government has approved Oxytec’s transgenic mosquitoes for commercial release in April 2014 [6].
In terms of safety, the new sex distortion transgenic mosquito is no better and possibly worse.
Homing endonuclease originally from slime mouldThe homing endonucleases are a collection of enzymes encoded either as freestanding genes located within introns (intervening noncoding sequence in protein-coding genes), as fusions with host proteins, or as self-splicing inteins (sequences within proteins that splice themselves out after translation). They cut genomic DNA within the cells that synthesize them at target sites. Repair of the cut DNA by the host cell frequently results in the gene encoding the homing endonuclease being copied into the cleavage site, hence the term 'homing' to describe the movement of these genes. Homing endonucleases can thereby transmit their genes horizontally within a host population, increasing their frequency more rapidly than classical Mendelian genes [7, 8].
Homing endonuclease (HEN) recognize target sequences of 15 to 40 bp long. Generally, owing to the homing mechanism, the gene encoding the endonuclease is located within the recognition sequence which the enzyme cuts, thus interrupting the homing endonuclease recognition sequence and limiting DNA cutting only to sites that do not (yet) carry the HEN. However, target recognition is not completely specific, and off target cleavage and integration has been known at least since the 1990s.
The I-Ppol homing endonuclease used in the construction of the sex distorter transgenic mosquitoes [1, 2] was originally isolated from the slime mould Physarum polycephalum. It is a member of the His-Cys box family found in myxomycetes and amoebae. The His-Cys box protein motif consists of two conserved histidine and three conserved cysteine residues within a 30 residue region of protein. These conserved His and Cys residues appear to contribute to a Zn2+-binding motif and to the endonuclease active site. The I-Ppol target site of 15 bp and cleavage position is as follows [9]. The cleavage at TTAA leaves a 4 nucleotide overhang at the cut ends.
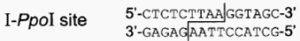
However, a substantial degree of degeneracy of the target sequence appeared to be tolerated, as demonstrated in a study in which partially randomized cleavage sites were created by mutagenesis [9]. Not only that, multiple target sites exist in all eukaryote genomes.
Target sites ubiquitous in all eukaryote species including humansThe native I-PpoI target site is by no means exclusive to the mosquito X-chromosome, it is actually present in multiples copies in the highly conserved 28S ribosome RNA (rRNA) genes of all eukaryotes species including humans, and currently being deployed in gene therapy experiments [10]. Each diploid human cell has about 600 copies of the rRNA genes in five clusters located to the short arms of chromosomes 13, 14, 15, 21, and 22. On account of the multitude of rRNA genes and the presence of possibly inert spacers between the gene repeats, the rDNA is considered an appealing safe haven for transgene integration. A team of researchers in Finland have created an HIV-1 integrase-1 PpoI fusion protein in order to guide the integration of an HIV viral vector into the rDNA [10].
The team also carried out a study on cytotoxicity of the construct. Although a genome-wide interaction study found that the IN-fusion PpoI proteins bind to their target sequence containing 28 S rRNA genes with 100-fold enrichment compared to controls, there was significant off-target binding to non-rRNA gene sites, and considerable cytotoxicity for all human cell lines tested from the double-stranded breaks in DNA produced by the I-PpoI endonuclease [11]. The researchers put a positive gloss on the findings by noting that the construct was more toxic to cancer than normal cells, and suggesting that the endonuclease could be used for cancer therapy.PiggBac a promiscuous transposon now used in gene therapy experimentsI have long warned against using promiscuous transposons as gene transfer vectors, especially for insects that bite people [12] (Terminator insects give wings to genome invaders, ISIS report). The piggyBac transposon was discovered in cell cultures of the moth Trichopulsia, the cabbage looper, where it caused high rates of mutations in the baculovirus infecting the cells by jumping into its genes [13] (Terminator insects –- a primer, ISIS Report). The piggyBac is 2.5kb long with 13 bp inverted terminal repeats. It has specificity for sites with the base sequence TTAA (same as the I-PpoI endonuclease cleavage site, see above). The probability of this sequence occurring at random in any genome is 0.254 or 0.4%. There is also evidence that the disabled piggyBac vector carrying the transgene, even when stripped down to the bare minimum of the border repeats, was nevertheless able to replicate and spread, because the transposase enzyme enabling the piggyBac inserts to move can be provided by transposons present in all genomes.
The main reason initially for using transposons as vectors in insect control was precisely because they can spread the transgenes rapidly by ‘non-Mendelian' means within a population, i.e., by replicating copies and jumping into genomes, thereby ‘driving’ the trait through the insect population. However, the scientists involved neglected the fact that the transposons could also jump into the genomes of the mammalian hosts including human beings. Although each transposon has its own specific transposase enzyme that recognizes its terminal repeats, the same enzyme can also interact with the terminal repeats of other transposons, and evidence suggests extensive cross-talk among related but distinct transposon families within a single eukaryotic genome (reviewed in [3]).
The use of the piggyBac transposon has been plagued by problems of instability in transformed Aedes aegypti [14]; and large unstable tandem inserts of the piggyBac transposon were prevalent [ 15].
In spite of instability and resulting genotoxicity, the piggyBac transposon has been used extensively also in human gene therapy [16]. Several human cell lines have been transformed, even primary human T cells using piggyBac [17]. These findings leave us little doubt that the transposon-borne transgenes in the transgenic mosquito can transfer horizontally to human cells. The piggyBac transposon was found to induce genome wide insertion mutations disrupting many gene functions.To concludeTransgenic mosquitoes are not the solution to eradicating dengue or malaria. On the contrary, they are among the most hazardous GMOs created, and should never be released into the wild on any commercial basis.Researchers should consider the risks involved before embarking on a project, and science journal editors and commentators should also question whether works for publications carry risks to health and the environment.
_______________
References1. “GM mosquitoes a ‘quantum leap’ towards tackling malaria”, Adam Vaughan, The Guardian, 10 June 2014,http://www.theguardian.com/environment/2014/jun/10/gm-mosquitos-malaria-genetic-modification
2. Galizi R, Doyle LA, Menishelli M, Bernardini F, Deredec A, Burt A, Stoddard BL, Windbichler N and Crisanti A. A synthetic sex-ratio distortion system for the control of the human malaria mosquito. Nature Communications 2014, published 10 June, doi:10.1038/ncommuns4977
3. Ho MW. Regulation of transgenic insects highly inadequate. Science in Society 54, 20-21, 2012.
4. Ho MW. Transgenic mosquitoes not a solution. Science in Society 54, 22-23, 2012.
5. Ho MW. Non-transgenic mosquitoes to control dengue. Science in Society 54, 24-25, 2012.
6. “Brazil approves use of genetically modified mosquitoes”, Hal Hodson, New Scientist, 23 April 2014,
http://www.newscientist.com/article/dn2 ... 6h_dHnjjIU7. Homing endonuclease, Wikipedia, 12 May 2014,
http://en.wikipedia.org/wiki/Homing_end ... d8928227-28. Belfort M and Perlman PS. Mechanisms of intron mobility. J Biol Chem 1995, 270, 30237-40.
9. Argast GM, Stephens KM, Emonds MJ and Monnat Jr RJ. I-PpoI and I-CreI homing site sequence degeneracy determined by random mutagenesis and sequential in vitro enrichment. J Mol Biol 1998, 280, 345-53.
10. Schenkwein D, Turkki V, Ahlroth MK, Timonen O, Airenne KJ and Ylä-Herttuaia S. rDNA-directed integration by an HIV-1 integrase-I-Ppol fusion protein. Nucleic Acids Research 2012, 1-10, doi:10.1093/nar/gks1438,http://nar.oxfordjournals.org/content/early/2012/12/25/nar.gks1438.full.pdf+html
11. Turkki V, Schenkwein D, Timonen O, Husso T, Lesch HP and Ylä-Herttuaia S. Lentiviral protein transduction with genome-modifying HIV-1 integrase-I-PpoI fusion proteins: studies on specificity and cytotoxicity. BioMed Research Interna 2014, article ID 370340, 11 pp,
http://www.hindawi.com/journals/bmri/2014/379340/12. Ho MW. Terminator insects give wings to genome invaders. ISIS Report, 19 March 2001,
http://www.i-sis.org.uk/terminsects-pr.php13. Cummins J. Terminator insects – a primer. ISIS Report, 15 March 2001,
http://www.i-sis.org.uk/piggybac-pr.php14. Adelman ZN1, Jasinskiene N1, Peek C1, Travanty EA2, Olson KE2, James AA1. Instability of the piggyBac element in transformedAedes aegypti. ISMIS 2002. Abstracts of the Fourth International Symposium on Molecular Insect Science. 70pp. Journal of Insect Science, 2, 17.
15. Adelman ZN, Jasinskiene N, Vally KJ, Peek C, Travanty EA, Olson KE, Brown SE, Stephens JL, Knudson DL, Coates CJ, James AA. Formation and loss of large, unstable tandem arrays of the piggyBac transposable element in the yellow fever mosquito, Aedes aegypti. Transgenic Res 2004, 13(5), 411-25.
16. Urschitz J, Kawasumi M, Owens J, Morozumi K, Yamashiro H, Stoytchev I, Marh J, Dee J Kawamoto K, Coates CJ, Kaminski JM, Pelczar P, Yanagimachi R, Moisyadi S. Helper-independent piggyBac plasmids for gene delivery approaches: strategies for avoiding potential genotoxic effects. Proc Natl Acad Sci U S A 2010, 107(18), 8117-22.
17. Galvan DL, Nakazawa Y, Kaja A, Kettlun C, Cooper LJ, Rooney CM, Wilson MH. Genome-wide mapping of PiggyBac transposon integrations in primary human T cells. J Immunother 2009, 32(8), 837-44.
18. Lobo N, Li X and Fraser Jr. MJ. Transposition of the piggyBac element in embryos of Drosophila melanogaster, Aedes aegypti andTrichoplusia ni. Mol Gen Genet 1999: 261: 803-10.
19. Fu Y, Foden JA, Khayter C, Maeder ML, Reyon D, Young JK and Sander JD. High frequency off-target mutagenesis induced by CRISPR-Cas nucleases in human cells. Nat Biotechnol 2013, 31, 822-6.
http://www.ncbi.nlm.nih.gov/pmc/article ... 488397.pdf