by Werner Slenczka and Hans Dieter Klenk
Institute for Virology, Philipps-Universität Marburg, Marburg, Germany
Reprints or correspondence: Dr. Hans Dieter Klenk, Institute for Virology, Philipps-Universität Marburg, Hans-Meerwein-Str. 2, 35043 Marburg, Germany ([email protected]).
NOTICE: THIS WORK MAY BE PROTECTED BY COPYRIGHT
YOU ARE REQUIRED TO READ THE COPYRIGHT NOTICE AT THIS LINK BEFORE YOU READ THE FOLLOWING WORK, THAT IS AVAILABLE SOLELY FOR PRIVATE STUDY, SCHOLARSHIP OR RESEARCH PURSUANT TO 17 U.S.C. SECTION 107 AND 108. IN THE EVENT THAT THE LIBRARY DETERMINES THAT UNLAWFUL COPYING OF THIS WORK HAS OCCURRED, THE LIBRARY HAS THE RIGHT TO BLOCK THE I.P. ADDRESS AT WHICH THE UNLAWFUL COPYING APPEARED TO HAVE OCCURRED. THANK YOU FOR RESPECTING THE RIGHTS OF COPYRIGHT OWNERS.
Abstract
Forty years ago, in early August 1967, the first filovirus ever detected, Marburg virus, made its appearance in Europe, causing severe and often fatal hemorrhagic fever in laboratory workers in Marburg and Frankfurt and, about 4 weeks later, in Belgrade. The etiological agent was isolated and identified by the combined efforts of virologists in Marburg and Hamburg within the very short time of 3 months. Marburg was not the only town whe re the virus was isolated and identified for the first time, but most cases of infection occurred in Marburg.
August 1967: Outbreak of a New Disease
In early August 1967, patients with unusual symptoms indicating an infectious disease were admitted to the university hospitals in Marburg and Frankfurt. The first patients were treated in their homes for up to 10 days, even though the illness was described as beginning suddenly with extreme malaise, myalgia, headache, and a rapid increase in temperature to as high as 39°C or more. Although the clinical symptoms were not very alarming during the first 3–4 days, additional symptoms and signs appeared at the end of the first week. Gastrointestinal symptoms, such as nausea, vomiting, and diarrhea, indicated to health care practitioners that the diagnosis might be dysentery or typhoid fever. The patients were therefore admitted to a hospital. At admission, most patients were observed to have conjunctivitis, exanthema, and enanthema, but shigellae or salmonellae were not found. During the second week after onset of disease, patient temperatures fell to 38°C, and petechiae and more-severe signs of hemorrhagic diathesis were recorded for ∼25% of patients. As indicated by transaminase levels, liver destruction reached its maximum at days 7 and 8 after onset of disease. Leukopenia with the appearance of immature polymorphonuclear leukocytes and thrombocytopenia (<10,000 cells/mm3) were detected. Patients were bleeding from all body orifices and from needle punctures. When the outcome was fatal, death occurred during the second week after onset of disease, at day 9 on average (range, day 7–16). In some cases, patients died from severe hemorrhagic shock on the day after hospital admission. Severe hemorrhagic signs, as seen in ∼25% of patients, were a signum mali ominis. All patients who died had hemorrhaging. Of 7 patients with manifest hemorrhages, 5 succumbed to the disease. Orchitis, a typical late-stage symptom, appeared in the third week after onset of disease or even at relapse during the fifth week. Mental confusion and paraesthesias were indicative of cerebral involvement. Relapses with hepatitis, orchitis, and uveitis with virus persisting in semen and in the anterior eye chamber were typical during the convalescent phase of both Marburg virus (MARV) and Ebola virus (EBOV) infections. In 1 case, a patient transmitted infection to his wife 120 days after onset of his disease, most probably by sexual intercourse. Virus was detectable in seminal fluid.
The incubation time of MARV disease could only be estimated retrospectively, after the source of infection and the date of exposure were known. Incubation ranged from 5 to 9 days, with an average of 8 days. The ratio of primary to secondary infections was 21:3 in Marburg, 4:2 in Frankfurt, and 1:1 in Belgrade. Three cases of secondary infection resulted from inadvertent needle-stick inoculations; in 1 case, a pathology technician cut himself on the forearm with a knife during a postmortem examination. Airborne transmission between humans did not occur, as indicated, for example, by the instance of a young man who slept in the same bed with his brother only a couple of days before he died; the brother did not develop disease and was seronegative for MARV 6 months later. One of the patients had been severely ill at the time of the outbreak but, for unknown reasons, was not hospitalized. He recovered and, 15 years later, maintained that he had had MARV disease. At that time, he underwent serological testing and was found to be seropositive for MARV antibody by IFA and ELISA. He had been exposed to monkey kidney cell cultures, which were used for the production of poliomyelitis vaccine. At 6 months after the outbreak, blood specimens were obtained from 120 persons who had been in close contact with patients or with infectious material but who had not developed disease. The specimens were tested for MARV antibody by complement fixation test, IFA, and ELISA and were found to be seronegative. Therefore, there was no indication of clinically inapparent infection.
While evaluating 32 patients at the 3 locations, we tried to estimate the influence of age and sex on the outcome of the disease. The mean duration of disease was 11.5 days in patients <20 years of age and reached 25.5 days in patients >40 years of age. In the younger age group, 33% of the patients died (n=9); in the older age group, the fatality rate was 20% (n=5). Although these data lack statistical significance, they suggest that the case fatality rate among older people is not higher than that among young patients. The influence of sex on outcome could not be assessed. There were 12 female patients: 2 of them died, but 4 of the survivors had cases of secondary infection, which, as indicated by general experience, results in milder disease symptoms, compared with cases of primary infection. Five of 24 male patients died, and 22 of these patients had cases of primary infection. Thus, the case fatality rate for cases of primary infection was 25% among female patients and 22.7% among male patients.
Three female patients became pregnant 1 or 2 years after infection. The pregnancy outcomes were normal in all 3 cases. The placentas were tested for virus and were negative. Umbilical cord blood was tested for antibody against MARV and was found to be positive for IgG but not for IgM antibody. After 12 months, the antibody titers had disappeared in the infants.
The Discovery of MARV
Microbiological and serological diagnostic studies covering most of the agents known at that time to cause hemorrhagic fever failed to reveal the etiology of the outbreak. When serological testing for leptospirosis was positive for some patients, Walter Mannheim, a bacteriologist in Marburg, tried to isolate leptospirae. On 22 August 1967, he inoculated 6 guinea pigs with 2-mL specimens of citrated blood, obtained from 3 patients on day 6 after onset of disease (specimens from each patient were given to 2 of the guinea pigs). The rectal temperatures of the 6 guinea pigs were taken on days 3, 4, and 5 postinoculation, and an increase in temperature to as high as 40°C or more was found in all the animals. However, in contrast with later animal passages of the agent, the guinea pigs did not show signs of severe disease. All animals were killed on 27 August, but leptospirae were not found in their blood.
Meanwhile, the high pathogenicity of the agent had become apparent: 5 patients had already died, and it therefore seemed to be too dangerous to continue the diagnostic work under very poor laboratory conditions. The blood specimens were stored in a freezer at −80°C. Specimens from the patients were transferred to national and international laboratories that had more expertise and were better equipped for work with such dangerous agents. The foreign laboratories were informed of the work that had already been done, including the results of the guinea pig inoculations. By mid September, it had become evident that the agent exhibited a low contagiousness. Only a few cases of secondary infection and no cases of tertiary infection had occurred, and no new cases had occurred during the previous 2 weeks. Therefore, Rudolf Siegert (figure 1) resumed experiments with guinea pigs, together with a Chinese colleague, Hsin Lu Shu. They found that the agent could be passed among guinea pigs and exhibited pathogenicity that increased from passage to passage. At the third passage, the animals fell ill with fever, hepatitis, and hemorrhagic disease that closely resembled human disease, and they died within 10 days after inoculation, with a marked drop in temperature. However, all efforts to determine the etiological agent by light or electron microscopy failed. Opportunistic bacterial infections were a major problem.

Figure 1. W.S. (left) and Rudolf Siegert (1914–1988; right) at the time of the initial outbreak of Marburg virus infection.
At this time, specimens of human and guinea pig convalescent serum were available, and some of the serum specimens were tagged with fluorescein for direct IFA. Three weeks later, W.S. (figure 1) detected intracytoplasmic inclusions in the tissues of infected guinea pigs, by IFA. Animals that had infected cells in the liver and spleen were selected for further studies using electronmicroscopy. Blood specimens from these animals were inactivated with formalin and sent to Dietrich Peters at the Bernhard Nocht Institute in Hamburg. Formalinized plasma was spun directly onto electron microscope (EM) grids, by means of a new technique developed by Gerhard Müller, and negative staining was done. By these methods, MARV was identified on 20 November, <3 months after the outbreak had begun (figure 2).
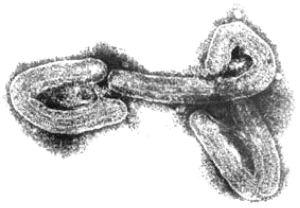
Figure 2. Electron micrograph of an isolate from the 1967 outbreak of Marburg virus infection.
The successful isolation of the virus and EM pictures of this etiologic agent, which exhibited a structure completely unknown at that time, were first reported to the scientific community at the Fourth Congreso Latinamericano de Microbiologia in Lima, Peru (26 November–2 December 1967), followed by publication in German in Deutsche Medizinische Wochenschrift on 22 December 1967 [1] and in English in German Medical Monthly on 1 January 1968 [2]. Antigen detection of MARV was also reported in 1968 [3]. Reprints of the original papers were sent to all the foreign institutions involved in the efforts to find the unknown agent of this new disease. Articles confirming the isolation and identification of the new virus were published by Kunz et al. [4] and by Kissling et al. [5] of the Centers for Disease Control and Prevention in 1968.
Where Did the Virus Come From?
Epidemiological studies were done in parallel with the microbiological studies. It became apparent very early that all the patients in Marburg were employees of Behringwerke, a producer of sera and vaccines, and that the patients in Frankfurt were employees of the Paul Ehrlich Institute, a control institute for sera and vaccines. The primary case patient in Belgrade, a veterinarian, was employed at Institute Torlak. A major activity of these institutions was the production and safety testing of live poliomyelitis vaccine. All patients with primary infection at the 3 locations had direct contact with blood, organs, and cell cultures from Cercopithecus aethiops monkeys. These animals were imported from Uganda and were used mainly for the production of kidney cell cultures, which were needed for the propagation of vaccine strains. Unfortunately, information on the health status of these monkeys is scarce and contradictory.
The fate and transportation route of the monkeys were complicated and only gradually revealed. Because of the Six Day War (5–10 June 1967), the monkey shipments from Uganda could not be transported directly to Frankfurt for distribution to their final destinations. Instead, they were brought to a London airport, where the airport employees were on strike at the time. Since no carrier was available for the transport from London to Frankfurt, the animals had to be kept in an animal house at the London airport, where they were caged in contact with finches from South America and langur monkeys from Ceylon (now Sri Lanka). In theory, the C. aethiops monkeys might have acquired an infectious agent from the finches or the langur monkeys, or they might have transmitted the virus to these animals. After a 2-day delay, the monkeys were finally transported, via the Frankfurt airport, to their final destinations in Belgrade, Frankfurt, and Marburg. It is clear that the 3 institutions received monkeys from the same shipments in June and July 1967. According to newspaper reports, 2 monkeys escaped from the shipment when the monkeys were transported to the animal house in London. These animals were found only a few days later and were shipped separately to Frankfurt. Fortunately, the monkeys did not distribute the virus in the London population.
Monkeys from 2 shipments were identified retrospectively as the most probable source of infection. These animals were received on 21 and 28 July 1967. Their health status seems to have been in the normal range, but they were killed soon after their arrival in Marburg and Frankfurt. An excess mortality rate of 33% was recorded only by the Institute Torlak in Belgrade, where the monkeys were kept for 6 weeks after arrival. However, whether the excess mortality was due to MARV infection was not proved. A peculiar procedure used by Ugandan monkey trappers might explain why MARV was imported to Europe by healthy looking animals. When the trappers found sick animals that could not be sold, they transported them to an island in Lake Victoria, leaving them there to die or to survive. When they did not have enough monkeys to complete a shipment, they rowed to this island, trapped some healthy looking animals, and included them in the shipment. Since cases of human infection have shown that MARV can persist for some months in sequestered parts of the body, such as the testes or anterior eye chambers, some healthy looking animals might have had MARV in their organs. At least 2 laboratory workers had acquired infection from contact with monkey kidney cell cultures.
MARV: 40 Years Later
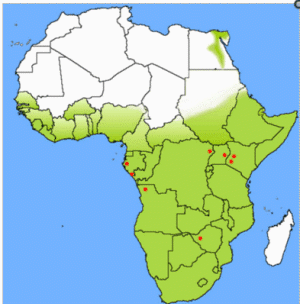
The assumption that MARV had originated on the African continent was very likely in 1967, but there was no direct proof of this hypothesis at that time, since the virus had not been identified in the incriminated monkeys. In addition, a small degree of uncertainty remained because of the contacts that the C. aethiops monkeys had while in the animal house in London.
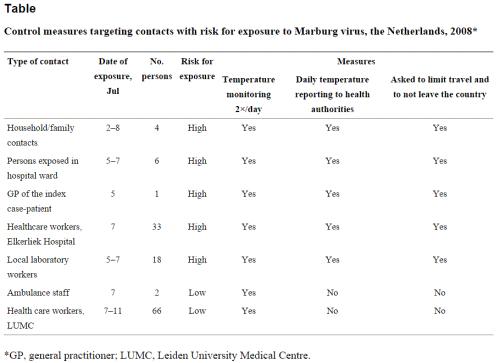
However, 8 years after the importation of MARV to Europe, a small outbreak of MARV disease occurred in South Africa. An Australian citizen had been hitchhiking in Rhodesia (now Zimbabwe) with a female companion. On their return to Johannesburg, he became ill with hemorrhagic symptoms and died a few days later in a hospital. His companion and a nurse developed a milder disease some days later, and both of them recovered. A strain of MARV that very closely resembled the strain from 1967 was isolated, and seroconversion to MARV was detected in the female patients during their convalescence. During his tour through Rhodesia, the index case patient had many contacts with wild animals but not with monkeys. In the following years, additional sporadic cases of MARV disease were observed (table 1). Except for a case that resulted from a laboratory accident in Russia, most of these infections occurred directly in or could be traced back to eastern Africa.
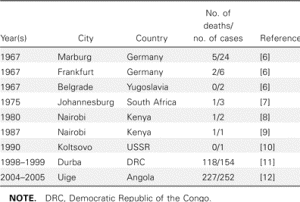
Table 1. Cases of Marburg virus infection, 1967–2005.
In contrast, when EBOV emerged in 1976, more-frequent and more-dramatic outbreaks were observed, with larger numbers of infected patients and higher case fatality rates. EBOV was soon found to be endemic in many countries of sub-Saharan Africa, ranging from the Ivory Coast to Sudan. Thus, in general, EBOV was thought to be more dangerous than MARV. However, the outbreaks of MARV infection in the Democratic Republic of the Congo in 1998–1999 and in Angola in 2004–2005 clearly indicated that this view had to be revised. Each outbreak resulted in nearly 200 deaths, and the mortality rate was similar to that attributed to Zaire EBOV (table 1). It is also evident that MARV is present in larger areas of Africa than had been previously acknowledged. Thus, MARV has to be considered as big a threat as EBOV.
MARV was discovered at a time when insight into the structure and replication of many viruses was being deepened enormously by the introduction of biochemical techniques. However, such studies were difficult to perform on MARV because of its high pathogenicity. A breakthrough came only 20 years later with the advent of recombinant DNA technology and the availability of appropriate biosafety containment procedures. The relationship to EBOV and the taxonomic assignment to the new filovirus family were established on a molecular level [13], and MARV has been shown to share gene structure, genome organization, and replication strategy with other members of the Mononegavirales order [14]. The reconstitution of a minigenome-based replication system [15] and the rescue of infectious virus from a full-length cDNA clone [16] will further promote our knowledge of the viral life cycle and pathogenesis. Alphavirus replicons expressing MARV proteins have elicited immune protection in cynomolgus monkeys [17], and the use of replicating vesicular stomatitis virus-based vectors have proved to be a particularly promising concept for vaccine development [18]. Thus, there is hope that, in the not-too-distant future, we will be able to control this deadly infection by immune prophylaxis and perhaps also by therapeutic measures. Ironically, the natural reservoir of the virus, the identification of which appeared to be a relatively easy task 40 years ago, is still a mystery.
Acknowledgments
Supplement sponsorship. This article was published as part of a supplement entitled “Filoviruses: Recent Advances and Future Challenges,” sponsored by the Public Health Agency of Canada, the National Institutes of Health, the Canadian Institutes of Health Research, Cangene, CUH2A, Smith Carter, Hemisphere Engineering, Crucell, and the International Centre for Infectious Diseases.
Footnotes
Potential conflicts of interest: none reported.
Financial support: Deutsche Forschungsgemeinschaft (grants SFB 286, 535, and 593). Supplement sponsorship is detailed in the Acknowledgments.
© 2007 by the Infectious Diseases Society of America
References
1. Siegert R, Shu HL, Slenczka W, Peters D, Mueller G. Zur Aetiologie einer unbekannten, von Affen ausgegangenen menschlichen Infektionskrankheit [abstract in English]. Dtsch Med Wochenschr (Stuttgart) 1967;92:2343. (Ger). 2370.
2. Siegert R, Shu HL, Slenczka HL, Peters D, Muller G. The aetiology of an unknown human infection transmitted by monkeys [preliminary communication]. Ger Med Mon 1968;13:1-2.
3. Slenczka W, Shu HL, Piepenburg G, Siegert R. Antigen-Nachweis des “Marburg-Virus” in den Organen infizierter Meerschweinchen durch Immunofluoreszenz [abstract in English]. Dtsch Med Wochenschr (Stuttgart) 1968;93:612-6. (Ger). 626.
4. Kunz C, Hofmann H, Kovac W, Stockinger L. Biologische und Morphologische Charakteristika des in Marburg aufgetretenen Hämorrhagischen Fiebers. Wien Klin Wochenschr 1968;80:161-2.
5. Kissling RE, Robinson RQ, Murphy FA, Whitfield SG. Agent of disease contracted from green monkeys. Science 1968;160:888-90.
6. Martini G, Siegert R, editors. Heidelberg and New York: Springer Verlag; 1971. Marburg virus disease.
7. Gear JS, Cassel GA, Gear AJ, et al. Outbreak of Marburg virus disease in Johannesburg. Br Med J 1975;4:489-93.
8. Smith DH, Johnson BK, Isaacson M, et al. Marburg-virus disease in Kenya. Lancet 1982;1:816-20.
9. Johnson ED, Johnson BK, Silverstein D, et al. Characterization of a new Marburg virus isolated from a 1987 fatal case in Kenya. Arch Virol Suppl 1996;11:101-14.
10. Nikiforov VV, Turovskii YI, Kalinin PP, et al. The laboratory case of Marburg hemorrhagic fever [in Russian]. Zurnal Mikrobiologii Epidemiologii Immunologii 1994;3:104-6.
11. Bausch DG, Nichol ST, Muyembe-Tamfum JJ, et al. Marburg hemorrhagic fever associated with multiple genetic lineages of virus. N Engl J Med 2006;355:909-19.
12. Towner JS, Khristova ML, Sealy TK, et al. Marburg virus genomics and association with a large hemorrhagic fever outbreak in Angola. J Virol 2006;80:6497-516.
13. Kiley MP, Cox NJ, Elliott LH, et al. Physicochemical properties of Marburg virus: evidence for three distinct virus strains and their relationship to Ebola virus. J Gen Virol 1988;69:1957-67.
14. Feldmann H, Mühlberger E, Randolf A, et al. Marburg virus, a filovirus: messenger RNAs, gene order, and regulatory elements of the replication cycle. Virus Res 1992;24:1-19.
15. Mühlberger E, Lötfering B, Klenk HD, Becker S. Three of the four nucleocapsid proteins of Marburg virus, NP, VP35, and L, are sufficient to mediate replication and transcription of Marburg virus-specific monocistronic minigenomes. J Virol 1998;72:8756-64.
16. Enterlein S, Volchkov V, Weik M, et al. Rescue of recombinant Marburg virus from cDNA is dependent on nucleocapsid protein VP30. J Virol 2006;80:1038-43.
17. Hevey M, Negley D, Pushko P, Smith J, Schmaljohn A. Marburg virus vaccines based upon alphavirus replicons protect guinea pigs and nonhuman primates. Virology 1998;251:28-37.
18. Jones SM, Feldmann H, Ströher U, et al. Live attenuated recombinant vaccine protects nonhuman primates against Ebola and Marburg viruses. Nat Med 2005;11:786-90.